Infectious Diseases
KEYWORDS
- Infection control
- Drug-resistant bacteria
- Hospital epidemiology
- Antimicrobial stewardship
- Real world data
- Viral infection
HEAD

LAB MEMBER
| Faculty | Position | Researchers |
|---|---|---|
| MORIOKA Hiroshi | Lecturer | Researchers |
| OKA Keisuke | Assistant Professor | Researchers |
| OKUMURA Toshihiko | Assistant Clinical Professor | Researchers |
CONTACT
| iga-ryu◎t.mail.nagoya-u.ac.jp (Please send a message after replacing "◎" mark with "@" mark. ) | |
| HP | Private Page |
OUTLINE
Our team currently consists of one professor, three instructors, and three medical staff members (including one graduate student).
Our basic research focuses on the epidemiology, infection control, and treatment of healthcare-associated infections and their causative pathogens, with a particular emphasis on multidrug-resistant Gram-negative bacilli. Additionally, we conduct surveillance and genomic typing of viral infections within Neonatal Intensive Care Units (NICU).
In terms of clinical research, we conduct multicenter cross-sectional epidemiological studies to promote hospital infection control and antimicrobial stewardship. We are currently engaged in multicenter surveys regarding outpatient antimicrobial prescription patterns, as well as the analysis of real-world data concerning perioperative antimicrobial use.
RESEARCH PROJECTS
This translation is written in a formal, academic style suitable for a research website, laboratory profile, or annual report.
1. Epidemiological Research on Healthcare-Associated Infections and Antimicrobial Prescribing
i) Cross-sectional Epidemiological Surveys of Inpatients
We introduced the ""Point Prevalence Survey (PPS)"" for infection control for the first time in Japan to investigate the status of healthcare-associated infections (HAIs) and antimicrobial use. Following a 2016 multi-center study across four university hospitals, we conducted a PPS involving 10,199 patients from 27 hospitals in Aichi Prefecture. The results revealed an HAI prevalence of 6.6% (device-associated infections at 0.91%), with 29.6% of all inpatients receiving antimicrobial agents. Furthermore, this study reported on the actual status of perioperative antimicrobial administration and the epidemiology of pediatric antimicrobial use in Aichi Prefecture. These findings represent the largest hospital epidemiological dataset in Japan and contribute to the improvement of infection control measures and antimicrobial stewardship. Moving forward, we plan to expand these PPS activities from Aichi Prefecture to a nationwide scale.
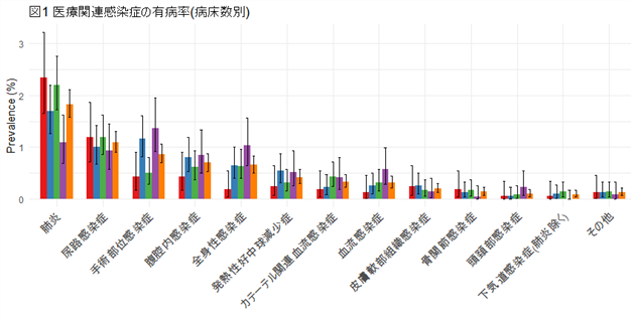
Prevalence of healthcare-associated infections by bed capacity.
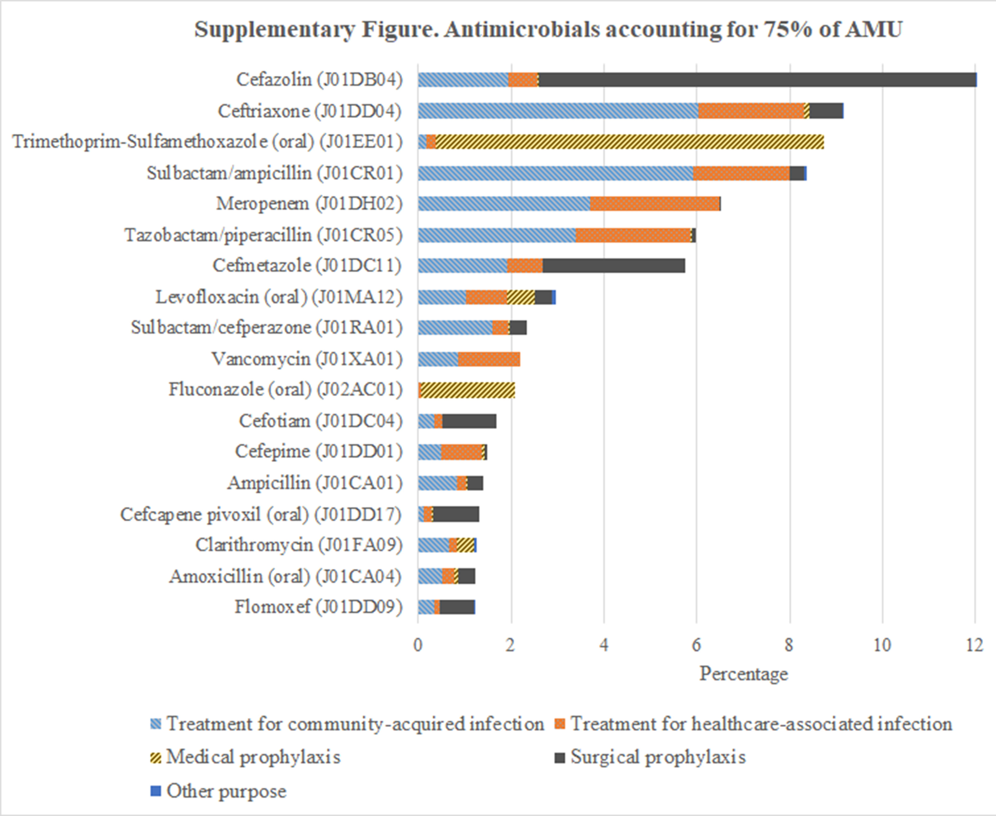
Drugs accounting for 75% of antimicrobial use and their indications.
ii) Research on Perioperative Antimicrobials
To identify issues in perioperative antimicrobial use, we surveyed 688 patients across 16 university hospitals in 2018. While 46.8% of patients overall received administration in accordance with guidelines, significant disparities in appropriate use were observed depending on the surgical specialty and facility. We are currently conducting research to evaluate the efficacy of perioperative antimicrobials using real-world data.
iii) Research on Outpatient Antimicrobial Prescribing
In fiscal year 2024, we conducted a survey of 2,987 individuals across 34 hospitals in Aichi Prefecture that have earned the ""Infection Control Enhancement Addition 1"" (Kansen-taisaku-kojo-kasan 1) to investigate inappropriate antimicrobial prescribing. Future plans include investigating the current status of outpatient antimicrobial administration using real-world data.
2. Analysis of Drug-Resistant Gram-Negative Bacilli
i) Analysis of Carbapenem-Resistant Enterobacterales (CRE)
The spread of CRE is becoming a global threat, and surveillance has commenced in Japan. Our laboratory conducts research on effective infection control measures and treatments for CRE, as well as molecular epidemiological analysis and drug-resistance mechanisms of CRE isolates detected in our hospital and regional surveillance.
We analyzed CRE isolates collected by the Council for Infection Control and Prevention at National and Public University Hospitals to identify factors associated with mortality and to analyze resistance genes and detected species.
We perform molecular epidemiological analysis on CRE strains where outbreaks are suspected within nearby facilities.
We utilize Next-Generation Sequencing (NGS) to analyze rare carbapenemase-producing organisms.
ii) Molecular Epidemiological Study of ESBL-Producing Klebsiella pneumoniae
Compared to E. coli, the isolation frequency of ESBL-producing Klebsiella pneumoniae is lower, and molecular epidemiological data remains scarce. In collaboration with Kyushu University and Asahikawa Medical University, our laboratory analyzed the ESBLs produced by these strains alongside their molecular and genetic characteristics. We found that the distribution of ESBL genes and quinolone resistance genes varied across the three geographic regions.
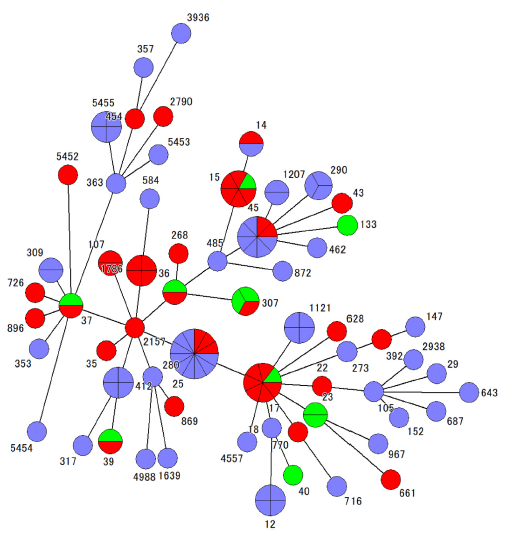
Cluster analysis regarding the distribution of ESBL-producing Klebsiella pneumoniae. Numbers indicate Sequence Types (ST). Circle size represents the number of isolates. Circle colors indicate the detecting university: Blue (Nagoya University Hospital), Green (Asahikawa Medical University Hospital), and Red (Kyushu University Hospital).
iii) Research on the Molecular Epidemiology, Resistance Mechanisms, and Clinical Presentation of the Enterobacter cloacae complex
The Enterobacter cloacae complex causes clinical conditions such as urinary tract and intra-abdominal infections, and resistance due to ESBL production or AmpC β-lactamase hyperexpression is a growing concern. As this complex consists of various subspecies, we collect and analyze isolates from multiple facilities to determine how resistance gene types, frequency of resistance, and clinical outcomes differ by subspecies.
3. Development of an Infectious Disease Management Support System (Joint Research with Shimadzu Corporation)
We developed a web-based communication system designed to streamline consultations regarding infectious disease clinical practice. By improving access to infectious disease specialists and contributing to earlier treatment decisions, the system aims to improve patient outcomes.
[Link: Development of Japan’s First ""Infectious Disease Management Support System"" through Industry-Academia Collaboration]
[Link: Interview: ""The Infectious Disease Management Support System Born from Field Needs is Launching Soon!""]
4. Research on Infection Control Derived from the Hospital Environment Focusing on Hand-Wash Sink Drains
To control infections originating from hospital hand-wash sinks, we are analyzing the microbiome on sink surfaces and within drainpipes. For bacterial contamination in drains, we are using the ""DrainPod,"" a drainpipe heating device developed by Moraine Corporation, to verify its effectiveness in reducing bacterial loads in existing piping, analyze deposits, and assess its impact on drain structure and safety. By combining culture methods, whole-genome sequencing, and endoscopic observation, we aim to establish infection control measures for the hospital environment that are grounded in clinical practice.
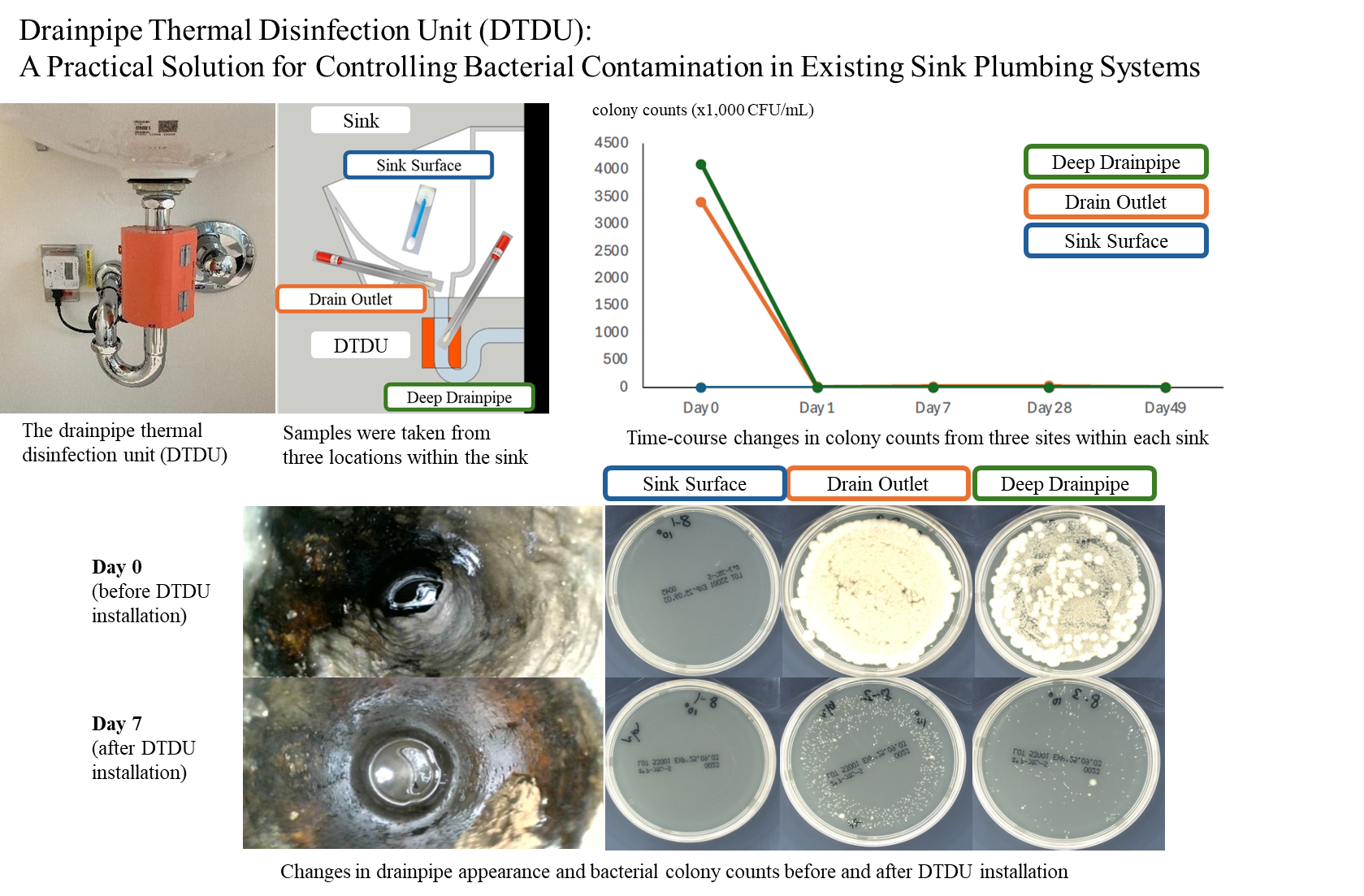
Verification of the disinfection effect of a drainpipe heating device on hospital hand-wash sink drains. It significantly reduced bacterial colony formation in existing pipes, and the antibacterial effect persisted with continuous use.
5. Research on Optimizing Alcohol-Based Hand Sanitizer Volume Using a Hand Hygiene Evaluation System
To improve the quality of hand hygiene in hospitals, we are conducting research using ""HandWe!"", a hand hygiene evaluation system developed by Moraine Corporation. In collaboration with infection control nurses, we are quantitatively evaluating the relationship between the volume of alcohol-based hand sanitizer, application coverage, and hand size to verify the optimal usage volume. We aim to establish practical, evidence-based hand hygiene techniques.
6. Research on Antibiograms for Medical Institutions and Clinics in Nagoya City
We are conducting a comparative analysis of microbial distribution and drug susceptibility using data from medical institutions in Nagoya with ""Infection Control Enhancement Addition 1"" and clinical data from the Falco Biosystems database. By comparing data from large-scale hospital groups with primary care clinics, we aim to construct antibiograms usable in primary care settings and establish optimal antimicrobial treatments tailored to real-world clinical practice.
7. Research on Optimizing Treatment for Bloodstream Infections Using Rapid Antimicrobial Susceptibility Testing (RAST)
We are conducting clinical microbiological research aimed at achieving rapid and appropriate antimicrobial therapy for bloodstream infections, which are severe infectious diseases. By establishing and operating RAST directly using positive blood culture samples in-hospital and validating it against the gold-standard broth microdilution method, we aim to optimize treatment rapidly. We plan to conduct multi-center joint research to build a system suited for the clinical landscape of bacteremia treatment in Japan.
8. Surveillance and Pathophysiological Analysis of Enterovirus/Parechovirus Infections in the Neonatal Intensive Care Unit (NICU)
While Enterovirus and Human Parechovirus infections are often mild or asymptomatic in children, they can become severe in neonates. However, identification of these virus species is not covered by insurance in Japan, making real-time clinical feedback difficult. We perform active surveillance using stool samples from patients in the NICU during epidemic periods via regular viral PCR assays. We also conduct genotyping to identify detected species and investigate factors involved in the progression to severe disease.
BIBLIOGRAPHY
2025
- Sahara S, Kinoshita T, Takimoto N, Oka K. A case of pyelonephritis and bacteremia caused by Candida glabrata in a patient on sodium glucose cotransporter 2 inhibitor, successfully treated with micafungin. J Pharm Health Care Sci. 2025 Aug 5;11(1):65.
- Kobayashi Y, Suzuki M, Umeda S, Oka K, Takahashi K, Shibayama K, Shibata S. Genomic insights into clinical non-O1/non-O139 Vibrio cholerae isolates in Japan. Microbiol Spectr. 2025 Aug 5;13(8):e0017525. Epub 2025 Jun 24.
- Morioka H, Ito K, Koizumi Y, Okudaira M, Tomita Y, Tsuji T, Akita K, Watariguchi T, Oka K, Watamoto K, Kato H, Ishihara M, Yokota M, Ito Y, Mutoh Y, Nagaoka M, Iwata S, Nozaki Y, Hamada H, Kojima Y, Kawasaki S, Hasegawa C, Shimizu J, Yagi T; Research Group of Aichi Point Prevalence Survey. Antimicrobial use in pediatric patients: a subgroup analysis of the 2020 point prevalence survey in Aichi, Japan. J Infect Chemother. 2025 Oct;31(10):102804. Epub 2025 Sep 2.
- Onishi K, Morioka H, Nishida K, Yamamoto M, Tsuchimoto D, Moriya Y, Kamihira O. Reducing infectious complications and healthcare costs in transrectal ultrasound-guided prostate biopsy with single-dose cefmetazole and levofloxacin. Urol Oncol. 2025 May;43(5):335.e1-335.e8. Epub 2025 Feb 11.
- Morioka H, Koizumi Y, Oka K, Okudaira T, Tomita Y, Kojima Y, Watariguchi T, Watamoto K, Mutoh Y, Tsuji T, Yokota M, Shimizu J, Hasegawa C, Iwata S, Nagaoka M, Ito Y, Kawasaki S, Kato H, Kitagawa Y, Hamada H, Nozak Y, Akita K, Shimizu S, Nozawa M, Kato M, Ishihara M, Ito K, Yagi T. Antimicrobial use in Japanese hospitals: results from a point-prevalence survey in Aichi, 2020. Journal of Hospital Infection. 2025 July, 161: 140-147.
- Hirao Y, Morioka H, Agata T, Iimura M, Taki S, Yagi T. Prolonged Bacillus cereus bacteremia: case report and literature review. J Infect Chemother. 2025 Jul;31(7):102736. Epub 2025 May 23.
- Hamada H, Morioka H, Okazaki M, Hashizume A, Kanda K, Oka K, Iguchi M, Yagi T. Re-evaluation of blood culture contamination rates: Discordance between clinical and laboratory assessment. J Infect Chemother. 2025 Apr;31(4):102628. Epub 2025 Jan 19.
- Osada Y, Oka K, Iguchi M, Morioka H, Iwata KI, Ohara M, Shimaoka N, Sawada T, Yagi T. Flavonifractor plautii bacteremia following bacterial translocation from the gut: A case report and literature review. J Infect Chemother. 2025 Mar;31(3):102592. Epub 2024 Dec 17.
- Ohashi K, Shinoda Y, Matsuoka T, Arai K, Hotta N, Takahashi T, Shikano H, Kagajo M, Yagi T, Usami E. Efficacy of Enhanced Antimicrobial Stewardship Team Interventions for Patients Receiving Meropenem and Tazobactam/Piperacillin. Biol Pharm Bull 2025;48(5):571-576.
- Park H, Nakamura N, Miyamoto S, Sato Y, Kim KS, Kitagawa K, Kobashi Y, Tani Y, Shimazu Y, Zhao T, Nishikawa Y, Omata F, Kawashima M, Abe T, Saito Y, Nonaka S, Takita M, Yamamoto C, Morioka H, Kato K, Sagou K, Yagi T, Kawamura T, Sugiyama A, Nakayama A, Kaneko Y, Shibata RY, Aihara K, Kodama T, Kamiyama A, Tamura T, Fukuhara T, Shibuya K, Suzuki T, Iwami S, Tsubokura M. Longitudinal antibody titers measured after COVID-19 mRNA vaccination can identify individuals at risk for subsequent infection. Sci Transl Med. 2025 Sep 17;17(816):eadv4214. Epub 2025 Sep 17.
2024
- Morioka H, Koizumi Y, Oka K, Okudaira M, Tomita Y, Kojima Y, Watariguchi T, Watamoto K, Mutoh Y, Tsuji T, Yokota M, Shimizu J, Hasegawa C, Iwata S, Nagaoka M, Ito Y, Kawasaki S, Kato H, Kitagawa Y, Goto T, Nozaki Y, Akita K, Shimizu S, Nozawa M, Kato M, Ishihara M, Ito K, Yagi T; Research Group of Aichi Point Prevalence Survey. Healthcare-associated infections in Japanese hospitals: results from a large-scale multicenter point-prevalence survey in Aichi, 2020. Infect Control Hosp Epidemiol. 2024 Oct 8:1-8. Online ahead of print.
- Sahara S, Kinoshita T, Amano T, Ishida M, Yamakita T, Takimoto N, Oka K. Decubitus ulcer infection and bacteremia due to tazobactam/piperacillin-resistant Veillonella parvula. Nagoya J Med Sci. 2024 Aug;86(3):524-530.
- Amano T, Nishikawa T, Oka K, Ota K, Shimizu T. How an Antimicrobial Stewardship Team Treated a Nocardia farcinica-Associated Brain Abscess: A Case Report. Cureus. 2024 Feb 21;16(2):e54605.
- Morita Y, Tanahashi K, Terashima-Murase C, Fukaura R, Oka K, Yagi T, Miyamoto Y, Ato M, Ishii N, Akiyama M. Mycobacterium marinum infection successfully treated with oral administration of minocycline and thermotherapy. Nagoya J Med Sci. 2024 Nov;86(4):699-702.
- Inagaki T, Asahi S, Ogawa K, Nakagawa T, Ohkura T, Osada Y, Nikai T, Yamada K, Yagi T, Uchiya K-i. Development of a rapid detection method for the macrolide resistance gene in Mycobacterium avium using the amplification refractory mutation system-loop-mediated isothermal amplification method. Microbiol Spectr. 2024 Apr 2;12(4):e0233923.
- Ito S, Muraki Y, Inose R, Mizuno K, Goto R, Kiyosuke M, Iinuma Y, Yagi T, Ohge H. Characteristics of pediatric patients claimed with acute upper respiratory infection during otorhinolaryngology consultations: A descriptive study of a large Japanese medical claims database. J Infect Chemother. 2024 Aug;30(8):815-819.
- Hirabayashi A, Yahara K, Oka K, Kajihara T, Ohkura T, Hosaka Y, Shibayama K, Sugai M, Yagi T. Comparison of disease and economic burden between MRSA infection and MRSA colonization in a university hospital: a retrospective data integration study. Antimicrob Resist Infect Control. 2024 Feb 29;13(1):27.
- Sarangi J, Ido A, Ito M, Iinuma C, Doyama Y, Jin W, Wachino J-i, Suzuki M, Iguchi M, Yagi T, Arakawa Y, Kimura K. Clinical isolates of Streptococcus mitis/oralis-related species with reduced carbapenem susceptibility, harboring amino acid substitutions in penicillin-binding proteins in Japan. Antimicrob Agents Chemother. 2024 Apr 3;68(4):e0117923.
- Morioka H, Koizumi Y, Watariguchi T, Oka K, Tomita Y, Kojima Y, Okudaira M, Ito Y, Shimizu J, Watamoto K, Kato H, Nagaoka M, Yokota M, Hasegawa C, Tsuji T, Shimizu S, Ito K, Kawasaki S, Akita K, Kitagawa Y, Mutoh Y, Ishihara M, Iwata S, Nozaki Y, Nozawa M, Kato M, Katayama M, Yagi T; Research Group of Aichi Point Prevalence Survey. Surgical antimicrobial prophylaxis in Japanese hospitals: Real status and challenges. J Infect Chemother. 2024 Jul;30(7):626-632.
- Isogai M, Kawamura K, Yagi T, Kayama S, Sugai M, Doi Y, Suzuki M. Evaluation of Klebsiella pneumoniae pathogenicity through holistic gene content analysis. Microb Genom. 2024 Sep;10(9).
- Kamiya S, Sugai T, Oka K, Noda E, Miyazaki A, Hagiwara-Fujishiro R, Yamada M, Yagi T, Akiyama M. Case of monomicrobial necrotizing fasciitis caused by extended-spectrum β-lactamase-producing Citrobacter freundii. J Dermatol. 2024 Apr 23.
- Fukuda Y, Morioka H, Yamamoto S, Iguchi M, Umeda S, Asahara T, Kanda K, Oka K, Nakayama G, Yagi T. Catheter-related bloodstream infection caused by Lacticaseibacillus paracasei: A case report and literature review. J Infect Chemother. 2024 Jul;30(7):664-667.
- Ohashi K, Matsuoka T, Shinoda Y, Takahashi T, Shikano H, Kagajo M, Yagi T, Usami E. Evaluation of long-term pharmacist-led prospective audit and feedback in antimicrobial stewardship: An 8-year study. Am J Infect Control. 2024 Jun;52(6):670-677.
- Hirao Y, Morioka H, Ozaki S, Kawachi M, Ban S, Yaguchi T, Watanabe A. A Case of Invasive Rhinosinusitis Caused by Penicillium brasilianum. Med Mycol J. 2024;65(4):111-115.
2023
- Fukaura R, Terashima-Murase C, Ota M, Noda H, Oka K, Ishihara Y, Shibayama K, Akiyama M. Methicillin-resistant Staphylococcus aureus-induced staphylococcal scalded skin syndrome in a preterm infant, and subsequent toxigenic analysis. J Dermatol. 2023 Dec;50(12):e422-e423. Epub 2023 Aug 22.
- Kobayashi H, Takeuchi S, Torii Y, Ikenouchi T, Kawada JI, Oka K, Kato S, Ogawa M. Time course of skin rash, computed tomography findings, and viral load in a rheumatoid arthritis patient with severe varicella pneumonia. IDCases. 2023 Jul 29;33:e01866.
- Oka K, Sahara S, Kuramae H, Osada Y. Mycobacterium obuense bacteremia: A case report and literature review. Int J Mycobacteriol. 2023 Jul-Sep;12(3):357-359.
- Kinoshita T, Sahara S, Amano T, Ito M, Sakakibara T, Takimoto N, Osada Y, Oka K. First Case Report of Peritoneal Dialysis-associated Peritonitis Caused by Lysinibacillus sphaericus. Intern Med. 2023 Oct 1;62(19):2919-2922. Epub 2023 Feb 22.
- Morioka H, Kaida H, Nishio M, Suga K. Peritonsillar abscess caused by Mycoplasma hominis and Fusobacterium necrophorum following oral sex. Auris Nasus Larynx. 2024 Apr;51(2):320-322. Epub 2023 Dec 2.
- Onishi K, Morioka H, Imaizumi T, Tsuchimoto D, Nishio M, Komiyama T. Risk factors for cefmetazole-non-susceptible bacteremia in acute cholangitis. J Infect Chemother. 2024 May;30(5):423-428. Epub 2023 Nov 18.
- Arisawa Y, Sugawara K, Morioka H, Takada K. Syphilis Showing Multiple Pulmonary Nodules without Respiratory Symptoms. Intern Med. 2024 Mar 15;63(6):885-886. Epub 2023 Aug 9.
- Aoki Y, Tamura T, Uchida W, Morioka H, Yamamoto M, Yuhara S, Nishio N, Mutsuga M, Furune S, Suzuki S, Nishiwaki K. Hypopharyngeal Injury by Transesophageal Echocardiography During Cardiac Surgery. J Cardiothorac Vasc Anesth. 2023 Oct;37(10):2027-2031. Epub 2023 Jun 19.
- Nishio M, Morioka H, Takai S, Osada Y, Seki Y, Osugi T, Oba A, Miyaki Y. Bacteraemia and obstructive pyelonephritis caused by Bifidobacterium breve in an elderly woman: a case report and literature review. Access Microbiol. 2023 Oct 19;5(10):000574.v3.
- Takamure K, Iwatani Y, Amano H, Yagi T, Uchiyama T. Inactivation characteristics of a 280 nm Deep-UV irradiation dose on aerosolized SARS-CoV-2. Environ Int. 2023; 177:108022.
- Kagami K, Ishiguro N, Iwasaki S, Usami T, Fukumoto T, Hayasaka K, Oyamada R, Watanabe T, Nakakubo S, Niinuma Y, Hagino T, Abe Y, Fujimoto I, Maekawa H, Fujibayashi R, Fuke S, Asahi K, Ota S, Nagakura T, Okubo T, Asanuma H, Ito T, Okano S, Komatsu E, Sasaki K, Hashimoto K, Washiya K, Kato Y, Kusumi K, Asai Y, Saito Y, Sakai Y, Sakurada M, Sakimoto Y, Ichikawa Y, Kinebuchi T, Kondo D, Kanno S, Kobayashi M, Hirabayashi K, Saitou S, Saito K, Ebina Y, Koshizaki Y, Chiba M, Yasuda A, Sato T, Togashi A, Abe T, Fujita T, Umehara K, Amishima M, Murakami N, Yagi T, Fujimoto S, Tajima T, Sugawara M, Takekuma Y. Correlation between antibiotic use and antibiotic resistance: A multicenter study using the Japan Surveillance for Infection Prevention and Healthcare Epidemiology (J-SIPHE) system in Hokkaido, Japan. Am J Infect Control. 2023 Feb;51(2):163-171.
2022
- Hara Y, Iguchi M, Tetsuka N, Morioka H, Hirabayashi A, Suzuki M, Tomita Y, Oka K, Yagi T. <Editors' Choice> Multicenter survey for carbapenemase-producing Enterobacterales in central Japan. Nagoya J Med Sci. 2022; 84(3):630-639.
- Sano M, Shindo Y, Takahashi K, Okumura J, Sakakibara T, Murakami Y, Iguchi M, Yagi T, Matsui S, Hasegawa Y. Risk factors for antibiotic resistance in hospital-acquired and ventilator-associated pneumonia. J Infect Chemother. 2022; 28(6):745-752.
- Oka K, Matsumoto A, Tetsuka N, Morioka H, Iguchi M, Ishiguro N, Nagamori T, Takahashi S, Saito N, Tokuda K, Igari H, Fujikura Y, Kato H, Kanai S, Kusama F, Iwasaki H, Furuhashi K, Baba H, Nagao M, Nakanishi M, Kasahara K, Kakeya H, Chikumi H, Ohge H, Azuma M, Tauchi H, Shimono N, Hamada Y, Takajo I, Nakata H, Kawamura H, Fujita J, Yagi T. Clinical characteristics and treatment outcomes of carbapenem-resistant Enterobacterales infections in Japan. J Glob Antimicrob Resist. 2022: 247-252.
- Sarangi J, Matsuo N, Nonogaki R, Hayashi M, Kawamura K, Suzuki M, Jin W, Tamai K, Ogawa M, Wachino JI, Kimura K, Yagi T, Arakawa Y. Molecular Epidemiology of Enterobacter cloacae Complex Isolates with Reduced Carbapenem Susceptibility Recovered by Blood Culture. Jpn J Infect Dis. 2022; 75(1):41-48
- Nonogaki R, Iijima A, Kawamura K, Kayama S, Sugai M, Yagi T, Arakawa Y, Doi Y, Suzuki M. PCR-based ORF typing of Klebsiella pneumoniae for rapid identification of global clones and transmission events. J Appl Microbiol. 2022; 133(3):2050-2062.
- Kobayashi H, Shindo Y, Kobayashi D, Sakakibara T, Murakami Y, Yagi M, Matsuura A, Sato K, Matsui K, Emoto R, Yagi T, Saka H, Matsui S, Hasegawa Y; Central Japan Lung Study Group. Extended-spectrum antibiotics for community-acquired pneumonia with a low risk for drug-resistant pathogens. Int J Infect Dis. 2022; 124:124-132.
- Nakai M, Oka K, Watanabe G, Kamei K, Tsukada N, Mori R, Nagaya M, Ukai Y, Tetsuka N, Morioka H, Iguchi M, Yagi T. Epidemiology and molecular characterization of fecal carriage of third-generation cephalosporin-resistant Enterobacterales among elderly residents in Japan. Journal of Infection and Chemotherapy. 2022; 28(4):569-575.
- Oka K, Tetsuka N, Morioka H, Iguchi M, Kawamura K, Hayashi K, Yanagiya T, Morokuma Y, Watari T, Kiyosuke M, Yagi T. Genetic and epidemiological analysis of ESBL-producing Klebsiella pneumoniae in three Japanese university hospitals. J Infect Chemother. 2022; 28(9):1286-1294.
- Morioka H, Ohge H, Nagao M, Kato H, Kokado R, Yamada K, Yamada T, Shimono N, Nukui Y, Yoshihara S, Sakamaki I, Nosaka K, Kubo Y, Kawamura H, Fujikura Y, Kitaura T, Sunakawa M, Yagi T. Appropriateness of surgical antimicrobial prophylaxis in Japanese university hospitals. J Hosp Infect. 2022; S0195-6701(22):00218-3.
- Murakami Y, Shindo Y, Sano M, Okumura J, Kobayashi H, Sakakibara T, Iguchi M, Takahashi K, Yagi T, Matsui S, Hasegawa Y. Effects of lymphocyte and neutrophil counts and their time courses on mortality in patients with postoperative pneumonia. Sci Rep. 2022 ;12(1):14564.
- Kobayashi Y, Shibata S, Yagi T. Molecular epidemiology and antimicrobial susceptibility profiles of Campylobacter jejuni isolated from bloodstream infections and enteritis in Japan. Diagnostic Microbiology and Infectious Disease. 2022; 103(2):115681.
- Morioka H, Oka K, Yamada Y, Nakane Y, Komiya H, Murase C, Iguchi M, Yagi T. Lysinibacillus fusiformis bacteremia: Case report and literature review. Journal of Infection and Chemotherapy 2022; 28(2):315-318.
- Sakakibara T, Shindo Y, Kobayashi D, Sano M, Okumura J, Murakami Y, Takahashi K, Matsui S, Yagi T, Saka H, Hasegawa Y. A prediction rule for severe adverse events in all inpatients with community-acquired pneumonia: a multicenter observational study. BMC Pulm Med. 2022; 22(1):34.
- Tetsuka N, Muramatsu H, Iguchi M, Oka K, Morioka H, Takahashi Y, Yagi T. Difficulties in diagnosing Malassezia furfur bloodstream infection and possibility of spontaneous resolution in a patient undergoing chemotherapy for neuroblastoma: A case report. J Infect Chemother. 2022; 28(7):987-990.
- Kagami K, Ishiguro N, Iwasaki S, Usami T, Fukumoto T, Hayasaka K, Oyamada R, Watanabe T, Nakakubo S, Niinuma Y, Hagino T, Abe Y, Fujimoto I, Maekawa H, Fujibayashi R, Fuke S, Asahi K, Ota S, Nagakura T, Okubo T, Asanuma H, Ito T, Okano S, Komatsu E, Sasaki K, Hashimoto K, Washiya K, Kato Y, Kusumi K, Asai Y, Saito Y, Sakai Y, Sakurada M, Sakimoto Y, Ichikawa Y, Kinebuchi T, Kondo D, Kanno S, Kobayashi M, Hirabayashi K, Saitou S, Saito K, Ebina Y, Koshizaki Y, Chiba M, Yasuda A, Sato T, Togashi A, Abe T, Fujita T, Umehara K, Amishima M, Murakami N, Yagi T, Fujimoto S, Tajima T, Sugawara M, Takekuma Y. Correlation between antibiotic use and antibiotic resistance: A multicenter study using the Japan Surveillance for Infection Prevention. Am J Infect Control. 2022; S0196-6553(22):00467-9.
- Akazawa N, Itoh N, Morioka H, Ogata T, Ishibana Y, Murakami H, Narita Y. Cholangitis with Sphingobacterium multivorum and Acinetobacter junii bacteremia in a patient with gastric cancer: A case report. J Infect Chemother. 2022 Oct;28(10):1419-1423. Epub 2022 Jun 16.
- Tsuchimoto D, Morioka H, Imaizumi T, Miyagawa S, Yamamoto M, Onishi K, Kuwabara Y, Takada K, Watamoto K. Current Status of Outpatient Oral Antimicrobial Prescription and the Influence of Antimicrobial Stewardship for Inpatients: A Repeated Cross-Sectional Study at a Japanese Community Hospital. Biol Pharm Bull. 2022;45(9):1340-1346.
- Kinoshita T, Sahara S, Mihara Y, Asai Y, Sato H, Sakakibara T, Takimoto N, Oka K. A case of acute focal bacterial nephritis caused by methicillin-resistant Staphylococcus saprophyticus in a 13-year-old adolescent girl treated with daptomycin. IDCases. 2022 Aug 6;29:e01594.
2021
- Sando Y, Morioka H, Sugawara K, Arisawa Y, Fukano H, Hoshino Y. A case of facial skin ulcer caused by Mycolicibacterium mageritense after tennis ball bruising. Int J Infect Dis. 2022 Jan;114:55-57. Epub 2021 Oct 27.
- Ryuge A, Saito S, Morioka H, Hachiya A, Kato N, Ishimoto T, Kosugi T, Maruyama S. Acquired Fanconi Syndrome in a Patient with Nontyphoidal Salmonella bacteremia. Intern Med. 2021 Mar 1;60(5):761-764. Epub 2020 Sep 30.
- Koshi E, Saito S, Okazaki M, Toyama Y, Ishimoto T, Kosugi T, Hiraiwa H, Jingushi N, Yamamoto T, Ozaki M, Goto Y, Numaguchi A, Miyagawa Y, Kato I, Tetsuka N, Yagi T, Maruyama S. Efficacy of favipiravir for an end stage renal disease patient on maintenance hemodialysis infected with novel coronavirus disease 2019. CEN Case Rep. 2021; 10(1):126-131.
- Morioka H, Iguchi M, Tetsuka N, Kinoshita F, Tomita Y, Kato D, Hirabayashi A, Matsumoto A, Oka K, Kato H, Inagaki T, Kato Y, Kitagawa K, Ichikawa K, Kouyama Y, Kawamura N, Toyotome Y, Adachi N, Ito Y, Yagi T. Five-year point prevalence survey of healthcare-associated infections and antimicrobial use in a Japanese university hospital. Infect Prev Pract. 2021; 3(3):100151
- Iwatsuki J, Kondo T, Takahashi N, Takami H, Nishigori H, Bustos-Villalobos I, Aleksic B, Kasuya H, Ban N, Yagi T, Skokauskas N. Problem-Based Learning in Child and Adolescent Psychiatry: A Perspective from Japan. Advances in Medical Education and Practice. 2021; 12: 1329-1335.
- Ebisui A, Inose R, Kusama Y, Koizumi R, Kawabe A, Ishii S, Goto R, Ishikane M, Yagi T, Ohmagari N, Muraki Y. Trends in Antipseudomonal Agent Use Based on the 2006 to 2015 Sales Data in Japan. Biological and Pharmaceutical Bulletin. 2021; 44(6):816-821.
- Oka K, Morioka H, Eguchi M, Sato Y, Tetsuka N, Iguchi M, Kanematsu T, Fukano H, Hoshino Y, Kiyoi H, Yagi T. Bursitis, Bacteremia, and Disseminated Infection of Mycobacteroides (Mycobacterium) abscessus subsp. Massiliense. Internal Medicine. 2021; 60(18):3041-3045.
2020
- Ishii S, Muraki Y, Kusama Y, Yagi T, Goto R, Ebisui A, Kawabe A, Inose R, Ohmagari N. The Trend for Antibiotic Use for Clostridioides (Clostridium) difficile Infection in Japan. Biol Pharm Bull. 2020; 43(4):693-696.
- Muraki Y, Kusama Y, Tanabe M, Hayakawa K, Gu Y, Ishikane M, Yamasaki D, Yagi T, Ohmagari N. Impact of antimicrobial stewardship fee on prescribing for Japanese pediatric patients with upper respiratory infections. BMC Health Serv Res. 2020; 20(1):399.
- Goto R, Inose R, Kusama Y, Kawabe A, Ishii S, Ebisui A, Ishikane M, Yagi T, Ohmagari N, Muraki Y. Trends of the Use of Anti-methicillin-Resistant Staphylococcus aureus Agents in Japan Based on Sales Data from 2006 to 2015. Biol Pharm Bull. 2020; 43(12):1906-1910.
- Ota A, Morita S, Matsuoka A, Shimaoka T, Maeda O, Mirsuma A, Yagi T, Asahara T, Ando Y. Detection of bacteria in blood circulation in patients receiving cancer chemotherapy. Int J Clin Oncol. 2020; 25(1):210-215.
- Sato Y, Iguchi M, Kato Y, Morioka H, Hirabayashi A, Tetsuka N, Tomita Y, Kato D, Yamada K, Kimura H, and Yagi T. Number of concomitant drugs with thrombocytopenic adverse effects and the extent inflammatory response resolution are risk factors for thrombocytopenia in patients treated with linezolid for more than 14 days. Nagoya J Med Sci. 2020; 82(3):407-414.
2019
- Tsutsui A, Yahara K, Clark A, Fujimoto K, Kawakami S, Chikumi H Iguchi M, Yagi T, Baker M A, O'Brien T, Stelling J. Automated detection of outbreaks of antimicrobial-resistant bacteria in Japan. The Journal of hospital infection. 2019; 102(2):226-233.
- Harada Y, Murata M, Matsumoto A, Kato D, Yagi T, Yaguchi T, Yoshikawa T, Takeichi T, Akiyama M, Yamaguchi Y, Koyama D, Terakura S, Nishida T, Kiyoi H. [Successful treatment of pre-engraftment disseminated fusariosis with high-dose liposomal]. Rinsho Ketsueki. 2019; 60(12):1641-1646.
- Tetsuka N, Hirabayashi A, Matsumoto A, Oka K, Hara Y, Morioka H, Iguchi M, Tomita Y, Suzuki M, Shibayama K, Yagi T. Molecular epidemiological analysis and risk factors for acquisition of carbapenemase-producing Enterobacter cloacae complex in a Japanese university hospital. Antimicrob Resist Infect Control. 2019; 8:126.
- Morioka H, Tokuda Y, Oshima H, Iguchi M, Tomita Y, Usui A, Yagi T. Fungal endocarditis after transcatheter aortic valve replacement (TAVR): Case report and review of literature. J Infect Chemother. 2019; 25(3):215-217.
2018
- Kato D, Morioka H, Tomita Y, Iguchi M, Hirabayashi A, Tetsuka N, Sadomoto T, Hyoudo M, Mochizuki M, Osada Y, Yamamoto M, Kato Y, Inagaki T, Ichikawa K, Yagi T. Active surveillance in response to the identification of a single carbapenemase-producing Escherichia coli at a Japanese university hospital. J Infect Chemother. 2018; 24(12):1013-1015.
- Morioka H, Nagao M, Yoshihara S, Ohge H, Kasahara K, Shigemoto N, Kajihara T, Mori M, Iguchi M, Tomita Y, Ichiyama S, Yagi T. The first multi-centre point-prevalence survey in four Japanese university hospitals. J Hosp Infect. 2018; S0195-6701(18):30143-9.
- Ohashi K, Matsuoka T, Shinoda Y, Fukami Y, Shindoh J, Yagi T, Yoshimura T, Sugiyama T. Evaluation of treatment outcomes of patients with MRSA bacteremia following antimicrobial stewardship programs with pharmacist intervention. Int J Clin Pract. 2018; 72(3):e13065.
- Horiba K, Kawada JI, Okuno Y, Tetsuka N, Suzuki T, Ando S, Kamiya Y, Torii Y, Yagi T, Takahashi Y, Ito Y. Comprehensive detection of pathogens in immunocompromised children with bloodstream infections by next-generation sequencing. Sci Rep. 2018; 8(1):3784.
- Sugawara G, Yokoyama Y, Ebata T, Igami T, Yamaguchi J, Mizuno T, Yagi T, Nagino M. Preoperative biliary colonization/infection caused by multidrug-resistant (MDR) pathogens in patients undergoing major hepatectomy with extrahepatic bile duct resection. Surgery. 2018; S0039-6060(18):30001-1.
- Sugawara G, Yokoyama Y, Ebata T, Mizuno T, Yagi T, Ando M, Nagino M. Duration of Antimicrobial Prophylaxis in Patients Undergoing Major Hepatectomy With Extrahepatic Bile Duct Resection: A Randomized Controlled Trial. Ann Surg. 2018; 267(1):142-148.
- Okumura J, Shindo Y, Takahashi K, Sano M, Sugino Y, Yagi T, Taniguchi H, Saka H, Matsui S, Hasegawa Y; Central Japan Lung Study Group. Mortality in patients with community-onset pneumonia at low risk of drug-resistant pathogens: Impact of β-lactam plus macrolide combination therapy. Respirology. 2018; 23(5):526-534.
- Yamasaki D, Tanabe M, Muraki Y, Kato G, Ohmagari N, Yagi T. The first report of Japanese antimicrobial use measured by national database based on health insurance claims data (2011-2013): comparison with sales data, and trend analysis stratified by antimicrobial category and age group. Infection. 2018; 46(2):207-214.
- Sugawara G, Yokoyama Y, Ebata T, Mizuno T, Yagi T, Ando M, Nagino M. Duration of Antimicrobial Prophylaxis in Patients Undergoing Major Hepatectomy With Extrahepatic Bile Duct Resection: A Randomized Controlled Trial. Ann Surg. 2018; 267(1):142-148.
2017
- Hirabayashi A, Kato D, Tomita Y, Iguchi M, Yamada K, Kouyama Y, Morioka H, Tetsuka N, Yagi T. Risk factors for and role of OprD protein in increasing minimal inhibitory concentrations of carbapenems in clinical isolates of Pseudomonas aeruginosa. J Med Microbiol. 2017; 66(11):1562-1572.
- Yamasaki D, Tanabe M, Muraki Y, Kato G, Ohmagari N, Yagi T. The first report of Japanese antimicrobial use measured by national database based on health insurance claims data (2011-2013): comparison with sales data, and trend analysis stratified by antimicrobial category and age group. Infection. 2017 Dec 22.
- Morioka H, Iguchi M, Kuzuya T, Mikamo H, Yagi T. Reccurent bacteremia and liver abscess caused by Clostridium difficile: A case report. Medicine (Baltimore). 2017; 96(35):e7969.
- Morioka H, Iguchi M, Oodate M, Yoneda M, Ushijima F, Hirabayashi A, Tetsuka N, Tomita Y, Kato D, Yagi T. Pneumococcal biliary tract infections-How rare are they? J Infect Chemother. 2017; 23(6):415-418.
- Kobayashi K, Imagama S, Kato D, Ando K, Hida T, Ito T, Tsushima M, Matsumoto A, Morozumi M, Tanaka S, Yagi T, Nishida Y, Ishiguro N. Collaboration with an infection control team for patients with infection after spine surgery. Am J Infect Control 2017; 45(7):767-770.
2016
- Adachi T, Ichikawa K, Inagaki T, Moriyama M, Nakagawa T, Ogawa K, Hasegawa Y, Yagi T. Molecular typing and genetic characterization of Mycobacterium avium subsp. hominissuis isolates from humans and swine in Japan. J Med Microbiol. 2016 Nov;65(11):1289-1295. Epub 2016 Sep 6.
- Muraki Y, Yagi T, Tsuji Y, Nishimura N, Tanabe M, Niwa T, Watanabe T, Fujimoto S, Takayama K, Murakami N, Okuda M. Japanese antimicrobial consumption surveillance: first report on oral and parenteral antimicrobial consumption in Japan (2009–2013). J Glob Antimicrob Resist. 2016 ;7:19-23
- Morioka H, Hirabayashi A, Iguchi M, Tomita Y, Kato D, Sato N, Hyodo M, Kawamura N, Sadomoto T, Ichikawa K, Inagaki T, Kato Y, Kouyama Y, Ito Y, Yagi T. The first point prevalence survey of health care-associated infection and antimicrobial use in a Japanese university hospital: A pilot study. Am J Infect Control 2016; 44(7):119-23.
- Morioka H, Yanagisawa N, Sasaki S, Sekiya N, Suganuma A, Imamura A, Ajisawa A, Kishida S. CD8 Encephalitis Caused by Persistently Detectable Drug-resistant HIV. Intern Med. 2016;55(10):1383-6. Epub 2016 May 15.
- Kato H, Yanagisawa N, Morioka H, Sasaki S, Sekiya N, Suganuma A, Imamura A, Ajisawa A. Laryngeal Kaposi's Sarcoma Complicated by the Immune Reconstitution Inflammatory Syndrome in an HIV-infected Patient. Intern Med. 2016;55(8):1001-5. Epub 2016 Apr 15.
2015
- Okachi S, Wakahara K, Kato D, Umeyama T, Yagi T, Hasegawa Y. Massive mediastinal cryptococcosis in a young immunocompetent patient. Respirol Case Rep 2015; 3(3):95-8.
- Ichikawa K, van Ingen J, Koh WJ, Wagner D, Salfinger M, Inagaki T, Uchiya K, Nakagawa T, Ogawa K, Yamada K, Yagi T. Genetic diversity of clinical Mycobacterium avium subsp. hominissuis and Mycobacterium intracellulare isolates causing pulmonary diseases recovered from different geographical regions. Infect Genet Evol 2015; 36:250-255.
- Shindo Y, Ito R, Kobayashi D, Ando M, Ichikawa M, Goto Y, Fukui Y, Iwaki M, Okumura J, Yamaguchi I, Yagi T, Tanikawa Y, Sugino Y, Shindoh J, Ogasawara T, Nomura F, Saka H, Yamamoto M, Taniguchi H, Suzuki R, Saito H, Kawamura T, Hasegawa Y; Central Japan Lung Study Group. Risk factors for 30-day mortality in patients with pneumonia who receive appropriate initial antibiotics: an observational cohort study. Lancet Infect Dis. 2015; 15(9):1055-1065.
- Uchiya K, Takahashi H, Nakagawa T, Yagi T, Moriyama M, Inagaki T, Ichikawa K, Nikai T, Ogawa K. Characterization lf a Novel Plasmid, pMAH135, from Mycobacterium avium Subsp. hominissuis. PLoS One 2015; 10(2):e0117797.
- Asai K, Yamada K, Yagi T, Baba H, Kawamura I, Ohta M. Effect of incubation atmosphere on the production and composition of staphylococcal biofilms. J Infect Chemother 2015; 21(1): 55-61.
- Ito R, Shindo Y, Kobayashi D, Ando M, Jin W, Wachino JI, Yamada K, Kimura K, Yagi T, Hasegawa Y, Arakawa Y. Molecular Epidemiological Characteristics of Klebsiella pneumonia Associated with Bacteremia among Patients with Pneumonia. J Clin Microbiol 2015; 53(3): 879-886.
2014
- Nakamura G, Wachino JI, Sato N, Kimura K, Yamada K, Jin W, Shibayama K, Yagi T, Kawamura K, Arakawa Y. Practical agar-based disk potentiation test for detection of fosfomycin-nonsusceptible Escherichia coli clinical isolates producing glutathione S-transferases. J Clin Microbiology 2014; 52(9):3175-3179.
2013
- Shindo Y, Ito R, Kobayashi D, Ando M, Ichikawa M, Shiraki A, Goto Y, Fukui Y, Iwaki M, Okamura J, Yamaguchi I, Yagi T, Tanikawa Y, Sugino Y, Shindou J, Ogasawara T, Nomura F, Saka H, Yamamoto M, Taniguchi H, Suzuki R, Saito H, Kawamura T, Hasagawa Y. Risk factors for drug-resistant pathogens in community-acquired and healthcare-associated pneumonia. Am J Respir Crit Care Med 2013;188(8):985-995.
- Yamaguchi M, Terao Y, Mori-Yamaguchi, Domon H, Sakaue Y, Yagi T, Nishino K, Yamaguchi A, Nizet V, Kawabata S. Streptococcus pneumoniae Invades Erythrocytes and Utilizes Them to Evade Human Innate Immunity. PLOS ONE. 2013; 8(10):e77282.
- Uchiya K, Takahashi H, Yagi T, Moriyama M, Inagaki T, Ichikawa K, Nakagawa T, Nikai T, Ogawa K. Comparative Genome Analysis of Mycobacterium avium Revealed Genetic Diversity in Strains that Cause Pulmonary and Disseminated Disease. PLOS ONE. 2013; 8(8):e71831
- Sugiura K, Sugiura N, Yagi T, Iguchi M, Ohno H, Miyazaki Y and Akiyama M. Cryptococcal cellulitis in a patient with bullous pemphigoid. Acta Derm Venereol. 2013; 93(2):187-188.
- Matsumura Y, Nagao M, Iguchi M, Yagi T, Komori T, Fujita N, Yamamoto M, Matsushima A, Takakura S, Ichiyama S. Molecular and clinical characterization of plasmid-mediated AmpC β-lactamase-producing Escherichia coli bacteraemia: a comparison with extended-spectrum β-lactamase-producing and non-resistant E. coli bacteraemia. Clin Microbiol Infect. 2013; 19(2):161-168.
MESSAGE
We welcome anyone interested in infectious disease research to contact us.

