Cancer Biology
KEYWORDS
- Epigenomics
- lncRNA
- Bacterial infection
- Glioblastoma
- Pancreas cancer
- Ovarian cancer
HEAD

LAB MEMBER
| Faculty | Position | Researchers |
|---|---|---|
| SHINJO Keiko | Lecturer | Researchers |
| SUZUKI Miho | Assistant Professor | Researchers |
| NISHIMURA Tatsunori | Assistant Professor | Researchers |
CONTACT
| iga-ryu◎t.mail.nagoya-u.ac.jp (Please send a message after replacing "◎" mark with "@" mark. ) | |
| HP | Private Page |
OUTLINE
In April 2017, I was appointed Professor of Cancer Biology in the Department of Oncology, Center for Neurological Diseases and Cancer, Graduate School of Medicine, Nagoya University. At this long-established and distinguished department, I am committed to further advancing medical research by building upon the strong scientific legacy cultivated by my predecessors, while also developing new and innovative research directions. Genes encoded in cellular DNA (the genome) must be precisely regulated so that they are selectively activated or silenced in response to cellular and environmental conditions. Epigenetics, which constitutes the central focus of our research, refers to the molecular mechanisms that determine how genetic information encoded in DNA is utilized without altering the DNA sequence itself. In this regulatory process, genes that are “active” or “inactive” are marked by specific molecular signatures. These signatures include various chemical modifications, such as DNA methylation, and are collectively referred to as the epigenome. In recent years, non-coding RNAs, which are not translated into proteins, have emerged as important regulators of epigenetic control and have attracted increasing attention. Recent advances in molecular biology and analytical technologies have led to remarkable progress in medical research. As a result, diseases for which the underlying mechanisms were previously unknown and effective treatments unavailable are now becoming amenable to therapeutic intervention. Such advances contribute directly to human health and represent a source of hope for society. By addressing unresolved medical challenges and approaching them from multiple scientific perspectives, we aim to elucidate the molecular mechanisms underlying disease pathogenesis and to develop novel diagnostic and therapeutic strategies.
RESEARCH PROJECTS
① Long non-coding RNA–mediated cancer pathogenesis
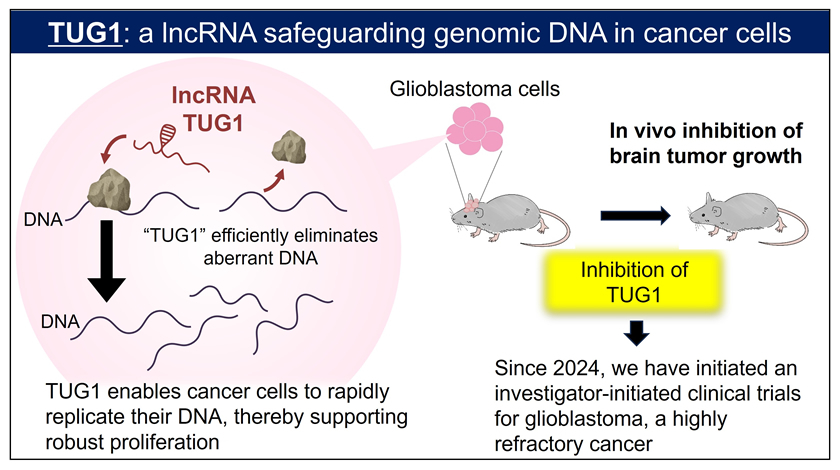
Because cancer cells actively divide and proliferate, they must replicate large amounts of DNA within a short period of time. We identified TUG1 (Taurine Upregulated Gene 1) as a long non-coding RNA that is rapidly produced in cells when problems occur during DNA replication. Our analyses revealed that, in cancer cells, TUG1 plays a crucial role in resolving abnormal DNA structures that interfere with DNA replication, thereby enabling replication to continue. This function is essential for cancer cell survival and is particularly prominent in glioblastoma (a malignant brain tumor), in which TUG1 is highly expressed. Indeed, inhibition of TUG1 using antisense oligonucleotides (TUG1-ASO) caused a marked arrest of DNA replication. As a result, irreparable DNA damage accumulated, ultimately leading to the induction of cell death. Based on these findings, in collaboration with the University of Tokyo, we developed a novel therapeutic agent, the antiTUG1, which combines TUG1-ASO with a drug delivery system (DDS) capable of efficiently delivering the agent to tumor sites. To evaluate the efficacy and safety of the antiTUG1 for glioblastoma in a clinical setting, an investigator-initiated clinical trial has been underway since February 2024 at Nagoya University Hospital, the National Cancer Center, and Kyoto University Hospital (see Bibliography 2025-2, 2024-2, 2023-1, 2021-1, 2019-2, 2017-1 and 2016-1).
In addition to TUG1, we are also conducting comprehensive studies on a wide range of long non-coding RNAs that possess diverse functions related to cancer malignancy and brain development.
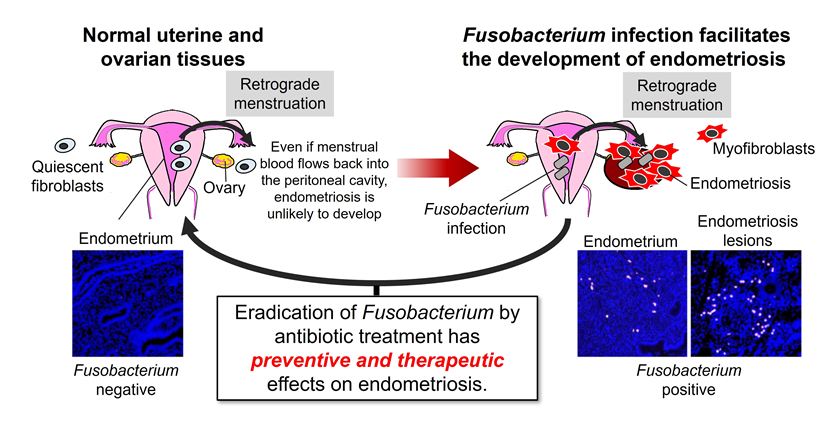
② Bacterial infection–mediated pathogenesis of precancerous lesions and cancer
Endometriosis, which forms lesions in the ovary, affects approximately 10% of women of reproductive age. It causes chronic pelvic pain and infertility and is also associated with an increased risk of malignant transformation in the future. Current treatments mainly rely on hormone therapy and surgery; however, these approaches have several limitations, including side effects, disease recurrence, and adverse effects on fertility. Therefore, the development of novel non-hormonal therapeutic strategies is strongly needed. We analyzed endometrial tissues from patients with endometriosis and discovered a high frequency of infection with the bacterium Fusobacterium. We found that Fusobacterium induces TGF-β production in endometrial tissues, which in turn increases the number of Transgelin (TAGLN)-positive fibroblasts, thereby promoting the development of endometriosis. In experiments using a mouse model of endometriosis, Fusobacterium infection exacerbated both the number and weight of lesions. In contrast, eradication of the bacteria using antimicrobial agents led to a marked improvement of the lesions. These findings indicate the potential of bacterial infection–targeted therapeutic approaches as a new treatment strategy. Furthermore, because endometriosis is considered a precancerous lesion for certain types of ovarian cancer, we are also investigating how Fusobacterium infection in endometriosis influences the initiation and malignant progression of ovarian cancer (see Bibliography 2023-2).
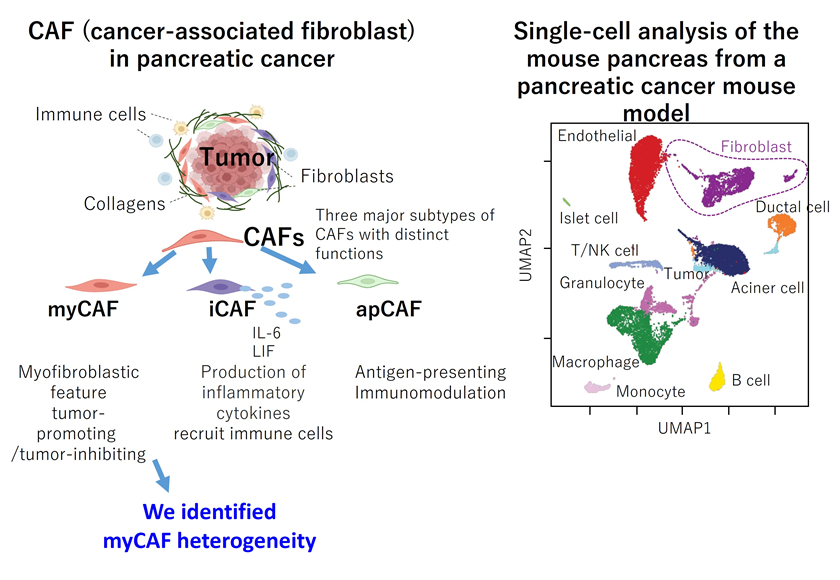
③ Fibroblast-mediated cancer pathogenesis
Fibroblasts are cells found in many organs, including the skin. Pancreatic ductal adenocarcinoma (PDAC) is characterized by a pronounced desmoplastic reaction, which is predominantly composed of cancer-associated fibroblasts (CAFs). Recent single-cell transcriptomic analyses of both mouse and human PDAC have identified three major subtypes of CAFs with distinct profiles: myCAF, iCAF, and apCAF. To delineate the epigenetic landscape and heterogeneity of CAFs in PDAC, we performed scATAC-seq analysis on pancreatic tissues from a PDAC mouse model. Epigenetic profiling revealed substantial heterogeneity within the myCAF population. Among myCAF subpopulations, Tagln emerged as a key functional mediator. Tagln knockout mice exhibited reduced collagen production and immune cell recruitment, suggesting that TAGLN-expressing myCAFs drive PDAC progression. These findings indicate that, in addition to targeting cancer cells themselves, therapeutic strategies aimed at stromal cells surrounding the tumor may represent an effective approach for pancreatic cancer treatment (see Bibliography 2025-1).
④ Cancer cells and epigenetics
The cells that make up our bodies originate from a single fertilized egg. Through repeated cell divisions, cells skillfully use (read) the same set of genes—our genetic “blueprint”—in different ways, resulting in the formation of diverse cell types such as skin cells and bone cells. The mechanisms that determine how genes are used are collectively referred to as epigenetics. The concept of epigenetics was proposed in 1942 by Conrad Waddington, even earlier than the DNA double-helix model proposed by Watson and Crick in 1953. Waddington defined epigenetics as the “causal mechanisms by which the genes of the genotype bring about phenotypic effects.” Because there is no exact equivalent term in Japanese, the word “epigenetics” is usually written in katakana. In China, it is translated as “表現遺伝学” (the study of expressed heredity), which may actually convey the meaning more clearly. Factors that define epigenetic regulation—including chromatin structure, histone modifications, DNA methylation, and non-coding RNAs—are collectively referred to as the epigenome, and under normal conditions, these factors are tightly regulated. However, aging and exposure to environmental factors can introduce chemical damage to these regulatory systems, leading to epigenomic abnormalities. It is thought that the accumulation of such abnormalities ultimately drives cellular transformation into cancer. Indeed, analyses of cancer cells have revealed that epigenomic abnormalities are detected in almost all cancers, influencing disease states from early-stage tumors to advanced malignancies. The fact that epigenomic abnormalities are present in all cancer cells suggests that their detection holds great promise for early cancer diagnosis. Moreover, unlike genetic mutations, epigenomic abnormalities are often reversible, meaning they can potentially be corrected by pharmacological interventions. Therefore, restoring normal epigenetic regulation in cancer cells represents a promising strategy for the development of novel cancer therapies. We are conducting in-depth studies of epigenetic abnormalities in cancer cells, aiming both to develop diagnostic approaches based on epigenetic changes and to create new therapeutic agents that normalize aberrant epigenetic states. Through these efforts, we seek not only to elucidate the mechanisms of carcinogenesis but also to advance cancer research toward direct clinical applications (see Bibliography 2024-1, 2022-1, 2020-2, 2019-1 and 2019-3).
BIBLIOGRAPHY
2025
- Shinjo K, Wang W, Kumegawa K, Maruyama R, Mii S, Shiraki Y, Tastunori Nishimura T, Murofushi Y, Suzuki M, Kabasawa T, Futakuchi M, Kanai A, Suzuki Y, Enomoto A, Kondo Y*. Transgelin Defines Pro-Tumorigenic Cancer-Associated Fibroblasts in Pancreatic Cancer. British Journal of Cancer. 2025, accepted.
- Deguchi S, Ohka F, Shiba Y, Yamaguchi J, Sato A, Shinjo K, Arakawa Y, Narita Y, Kondo Y, Saito R. Investigator-initiated phase I trial of an oligonucleotide therapeutic targeting long noncoding RNA TUG 1 for recurrent glioblastoma. BMC Cancer. 2025, 25(1):251.
2024
- Shinjo K, Umehara T, Niwa H, Sato S, Katsushima K, Sato S, Wang X, Murofushi Y, Suzuki MM, Koyama H, Kondo Y*. Novel pharmacologic inhibition of lysine-specific demethylase 1 as a potential therapeutic for glioblastoma. Cancer Gene Ther. 2024, 31(12):1884-1894.
- Sonobe R, Yang P, Suzuki MM, Shinjo K, Iijima K, Nishiyama N, Miyata K, Kataoka K, Kajiyama H, Kondo Y*. Long noncoding RNA TUG1 promotes cisplatin resistance in ovarian cancer via upregulation of DNA polymerase eta. Cancer Sci. 2024, 115(6): 1910-1923.
2023
- Suzuki MM, Iijima K, Ogami K, Shinjo K, Murofushi Y, Xie J, Wang X, Kitano Y, Mamiya A, Kibe Y, Nishimura T, Ohka F, Saito R, Sato S, Kobayashi J, Yao R, Miyata K, Kataoka K, Suzuki HI, Kondo Y*. TUG1-mediated R-loop resolution at microsatellite loci as a prerequisite for cancer cell proliferation. Nat Commun. 2023, 14(1):4521.
- Muraoka A, Suzuki M, Hamaguchi T, Watanabe S, Iijima K , Murofushi Y ,Shinjo K, Osuka S, Hariyama Y, Ito M, Ohno K, Kiyono T, Kyo S, Iwase A, Kikkawa F, Kajiyama H, Kondo Y*. Fusobacterium infection facilitates the development of endometriosis through the phenotypic transition of endometrial fibroblasts. Sci Transl Med. 2023, 15(700):eadd1531.
2022
- Matsumoto Y, Shinjo K, Mase S, Fukuyo M, Aoki K, Ozawa F, Yoshihara H, Goto S, Kitaori T, Ozaki Y, Takahashi S, Kaneda A, Sugiura-Ogasawara M, Kondo Y*. Characteristic DNA methylation profiles of chorionic villi in recurrent miscarriage. Sci Rep. 2022, 12(1):11673.
2021
- Tasaki T, Suzuki M, Katsushima K, Shinjo K, Iijima K, Murofushi Y, Naiki-Ito A, Hayashi K, Qiu C, Takahashi A, Tanaka Y, Kawaguchi T, Sugawara M, Kataoka T, Taito M, Miyata K, Kataoka K, Noda T, Gao W, Kataoka H, Takahashi S, Kimura K, Kondo Y*. Cancer-specific targeting of taurine upregulated gene 1 enhances the effects of chemotherapy in pancreatic cancer. Cancer Res. 2021, 81(7):1654-1666.
- Yamazaki S, Ohka F, Hirano M, Shiraki Y, Motomura K, Tanahashi K, Tsujiuchi T, Motomura A, Aoki K, Shinjo K, Murofushi Y, Kitano Y, Maeda S, Kato A, Shimizu H, Yamaguchi J, Adilijiang A, Wakabayashi T, Saito R, Enomoto A, Kondo Y, Natsume A. Newly established patient-derived organoid model of intracranial meningioma. Neuro Oncol. 2021, 23:1936-1948.
2020
- Totani H, Shinjo K, Suzuki M, Katsushima K, Mase S, Masaki A, Ito A, Ri M, Kusumoto S, Komatsu H, Ishida T, Inagaki H, Iida S, Kondo Y*. Autocrine HGF/c-Met signaling pathway confers aggressiveness in lymph node adult T-cell leukemia/lymphoma. Oncogene. 2020, 39:5782-5794.
- Shinjo K, Hara K, Nagae G, Umeda T, Katsushima K, Suzuki M, Murofushi Y, Umezu Y, Takeuchi I, Takahashi S, Okuno Y, Matsuo K, Ito H, Tajima S, Aburatani H, Yamao K, Kondo Y*. A Novel Sensitive Detection Method for DNA Methylation in Circulating Free DNA of Pancreatic Cancer. PLoS One. 2020, 15:e0233782.
2019
- Ohka F, Shinjo K, Deguchi S, Matsui Y, Okuno Y, Katsushima K, Suzuki M, Kato A, Ogino N, Yamamichi A, Aoki K, Suzuki H, Sato S, Arul Rayan N, Prabhakar S, Göke J, Shimamura T, Maruyama R, Takahashi S, SUzumura A, Kimura H, Wakabayashi T, Zong H, Natsume A, Kondo Y*. Pathogenic Epigenetic Consequences of Genetic Alterations in IDH-Wild-Type Diffuse Astrocytic Gliomas. Cancer Res. 2019, 79:4814-4827.
- Watanabe S, Hayashi K, Toh K, Kim HJ, Liu X, Chaya H, Fukushima S, Katsushima K, Kondo Y, Uchida S, Ogura S, Nomoto T, Takemoto H, Cabral H, Kinoh H, Tanaka HY, Kano MR, Matsumoto Y, Fukuhara H, Uchida S, Nangaku M, Osada K, Nishiyama N, Miyata K, Kataoka K. In vivo rendezvous of small nucleic acid drugs with charge-matched block catiomers to target cancers. Nat Commun. 2019, 10:1894.
- Mase S, Shinjo K, Totani H, Katsushima K, Arakawa A, Takahashi S, Lai HC, Lin RI, Chan MWY, Sugiura-Ogasawara M, Kondo Y*. ZNF671 DNA methylation as a molecular predictor for the early recurrence of serous ovarian cancer. Cancer Sci. 2019, 110:1105-1116.
2017
- Deguchi S, Katsushima K, Hatanaka A, Shinjo K, Ohka F, Wakabayashi T, Zong H, Natsume A, Kondo Y*. Oncogenic effects of evolutionarily conserved noncoding RNA ECONEXIN on gliomagenesis. Oncogene. 2017, 36:4629-4640.
2016
- Katsushima K, Natsume A, Ohka F, Shinjo K, Hatanaka A, Ichimura N, Sato S, Takahashi S, Kimura H, Totoki Y, Shibata T, Naito M, Kim HJ, Miyata K, Kataoka K, Kondo Y*. Targeting the Notch-regulated non-coding RNA TUG1 for glioma treatment. Nat Commun. 2016, 7:13616.
MESSAGE
Engaging in basic research is not something that is enjoyable every single day. Nevertheless, we firmly believe that research accumulated through steady and persistent efforts will one day help save patients suffering from intractable diseases.
For those who are interested in the atmosphere of our laboratory or our research themes, we warmly invite you to visit us and feel free to contact us for a consultation. You are most welcome to stop by simply to listen or to ask questions. Please do not hesitate to visit us.

